Abstract
Background:
Genetic information has become critical to understand the development of T-cell acute lymphoblastic leukemia (T-ALL) and to elucidate the origin of disease relapse. Several genetic markers, together with measurable residual disease (MRD), are considered strong predictors of patient outcome. However, the prognostic significance of genetic markers can varie according to treatment.
Aim: We used targeted deep sequencing to analyze the genetic profile of 125 T-ALL patients enrolled in three consecutive MRD-oriented trials from the Spanish PETHEMA (Programa Español de Tratamientos en Hematología) group. Genomic information was analyzed together with the main clinical and biologic data in a subset of 111 patients with detailed clinical and outcome data to determine the prognostic significance for overall survival (OS) and cumulative incidence of relapse (CIR).
Methods: Genetic mutations were detected using a custom gene panel and sequenced on a MiSeq platform. Alignment, variant calling, filtration and annotation of variants were done using standardized pipelines. OS curves were plotted by the Kaplan-Meier method and compared by the log-rank test. CIR was estimated using cumulative incidence functions by competing risks analysis. A Cox proportional hazard regression model was used to identify predictive factors for OS. Statistical significance was set at (two-sided) p-values <0.05.
Results: Recurrently mutated genes found in ≥4/125 patients involved transcription factor tumor suppressor genes (PTEN, BCL11B, RUNX1, GATA3, ETV6), epigenetic regulators (PHF6, DNMT3A, EP300, KMT2C, EZH2, TET2), DNA mismatch repair genes (MSH2), ribosomal (RPL5) and RNA splicing (U2AF1) genes, and genes involved in the RAS/MAPK (NRAS), WNT (FAT1, FAT3), IL7R-JAK-STAT (JAK3, JAK1, IL7R) and NOTCH1 signaling pathways, respectively. Mutations in the latest pathway (NOTCH1 & FBXW7) was found in 88/125 (70%) patients.
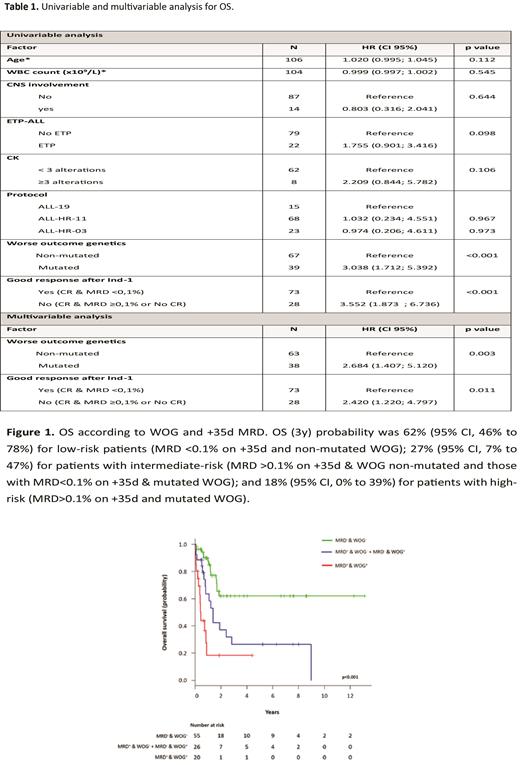
Clinical-genetic correlations revealed that patients with mutations in JAK3, DNMT3A, N/KRAS, IL7R, MSH2 or in U2AF1 were associated with lower OS (vs unmutated patients). None of the mutated genes had impact on CIR. Upon grouping the mutated genes according to their functional role and potential biological impact on T-ALL, two gene signatures were defined. These included the aging gene signature (DNMT3A and U2AF1) characterized by mutations in genes identified in clonal hematopoiesis of indeterminate potential (CHIP); and the treatment resistance gene signature (JAK3, N/KRAS, IL7R and MSH2), defined by mutations in genes involved in resistance to the ALL therapy. Both clusters identified patients with poorer response to therapy (poorer blast clearance on day 14 of induction treatment and lower CR rates). Therefore, we considered together (worse outcome genetics [WOG] signature) for univariate and multivariate analyses. WOG and MRD level (0.1% cut-off) on day 35 after induction therapy (+35d MRD) showed significant prognostic impact in the univariable and multivariable analyses for OS (3y) with a hazard ratio (95% CI) of 2.4 (1.2; 4.8) and 2.7 (1.4; 5.1), respectively (Table 1). OS according to these two variables allowed risk stratification of T-ALL into low, intermediate- and high-risk (HR) patients with significantly different outcomes (p<0.001) (Figure 1).
Conclusion: A genetic signature with independent prognostic significance of MRD has been identified in this cohort of patients included in MRD-oriented trials. This gene signature (WOG) together with MRD could help to improve risk-stratification of adult T-ALL patients and would be of interest in the search for new therapies for HR patients
Funding: Support from AECC (GC16173697BIGA); ISCIII (PI19/01828 and PI19/01183), co-funded by ERDF/ESF, "A way to make Europe"/"Investing in your future", CERCA/Generalitat de Catalunya SGR 2017 288 (GRC)/ C González-Gil was supported by AGAUR grant (2020 FI_B2 00210).
Diaz-Beyá: Jazz: Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Astellas: Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Novartis: Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Celgene: Membership on an entity's Board of Directors or advisory committees, Speakers Bureau. Mercadal: Gilead Sciences, Inc.: Honoraria, Speakers Bureau; Roche: Honoraria, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau. Tormo: Astellas: Consultancy, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Novartis: Consultancy, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Jazz Pharmaceuticals: Consultancy, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Pfizer: Consultancy, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau; Amgen: Consultancy, Membership on an entity's Board of Directors or advisory committees, Speakers Bureau. Barba: Amgen, Celgene, Gilead, Incyte, Jazz Pharmaceuticals, MSD, Novartis, Pfizer and Roche, Jazz Phar,aceuticals: Honoraria; Cqrlos III heqlth Institute, aSOCIACION espanola contra el cancer, PERIS: Research Funding. Maciejewski: Regeneron: Consultancy; Novartis: Consultancy; Bristol Myers Squibb/Celgene: Consultancy; Alexion: Consultancy. Ribera: ARIAD: Consultancy, Research Funding, Speakers Bureau; AMGEN: Consultancy, Research Funding, Speakers Bureau; Pfizer: Consultancy, Research Funding, Speakers Bureau; TAKEDA: Consultancy, Research Funding, Speakers Bureau; NOVARTIS: Consultancy, Speakers Bureau; SHIRE: Consultancy, Speakers Bureau.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal